|
This script takes w_trace output as a command line argument, loads the format ( float ( frame ), float ( frame ), float ( frame ))) File ( 'west.h5' ) coords = for iteration, seg_id in infile : iter_key = "iter_ \n ". 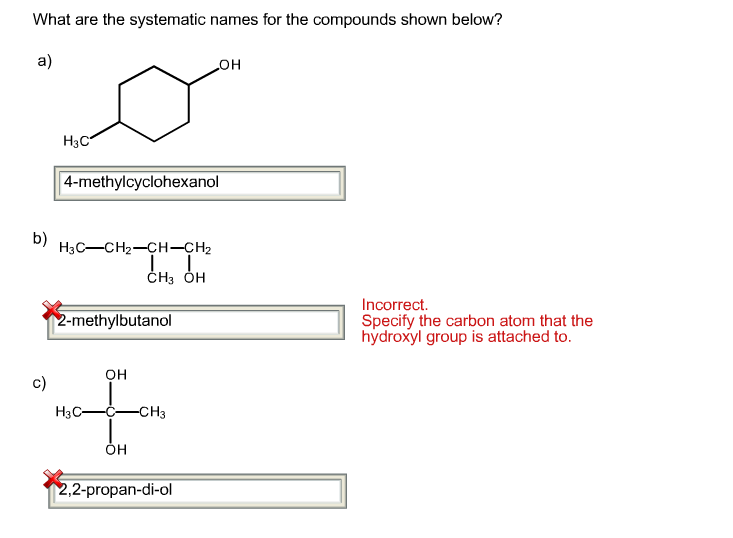
Import h5py, numpy, sys infile = numpy. NAMD configuration files support tcl scripting, which we use to export eachįrame of the trajectory after dynamics have been run. Simulations, may cause problems as simulations started simultaneously from the Setting the random seed based on the time, as is common for brute-force Is also necessary to set the random seed to a different value for each segment. 
Thermostat WESTPA requires a stochastic element such that trajectoriesīranched from a common point diverge. Second is the use of the stochastic Langevin The appropriate τ depends on the system and progress coordinate, and for The the segment duration, τ, which is equal to the timestep times the number of Several settings in this configuration file are important to WESTPA. dcd # Trajectory outfile dcdfreq 50 # Trajectory output interval (timesteps) # INTEGRATOR # timestep 2.0 # Simulation timestep (fs) run 250 # Simulation duration (timesteps) prm # Parameter infile seed RAND # Use random seed from WESTPA # ENSEMBLE # langevin on # Langevin thermostat enabled temperature 300 # Initial temperature (K) langevindamping 0.5 # Thermostat damping constant (ps -1) # NONBONDED INTERACTIONS # cutoff 999.0 # Nonbonded cutoff exclude scaled1 - 4 # Scaling between 1-4 bonded atoms 1 - 4 scaling 1.0 # 1-4 interaction scaling switching off # Van der Waals switching disabled pairlistdist 1000.0 # Neighbor list cut-off (A) stepspercycle 250 # Neighbor list update interval (timesteps) # IMPLICIT SOLVENT # GBIS on # Generalized Born implicit solvent # OUTPUT # outputname seg # Output file prefix outputEnergies 50 # Energy log output interval (timesteps) restartfreq 0 # Restart file output interval (timesteps) binaryoutput no # Do not output binary trajectory dcdfile seg. 
psf # Topology infile parameters par_all27_prot_na. coor # Structure infile velocities parent. # 0.5 ps NVT production with Langevin thermostat and GB implicit solvent # INPUT # paraTypeCharmm on # Parameter infile in 'CHARMM' format coordinates parent.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
